x
This website uses cookies to improve user experience. By using NMPFamDB you consent to all cookies in accordance with our
privacy policy.
OK
Sample 3300030719
3300030719: Metatranscriptome of soil microbial communities from Risofladan, Vaasa, Finland - UN-2 (Metagenome Metatranscriptome)
Overview
| Basic Information |
|---|
| IMG/M Taxon OID | 3300030719 Open in IMG/M |
GOLD Reference
(Study | Sequencing Project | Analysis Project) | Gs0135150 | Gp0323847 | Ga0307926 |
| Sample Name | Metatranscriptome of soil microbial communities from Risofladan, Vaasa, Finland - UN-2 (Metagenome Metatranscriptome) |
| Sequencing Status | Permanent Draft |
| Sequencing Center | DOE Joint Genome Institute (JGI) |
| Published? | N |
| Use Policy | Open |
| Dataset Contents |
|---|
| Total Genome Size | 51523379 |
| Sequencing Scaffolds | 6 |
| Novel Protein Genes | 8 |
| Associated Families | 7 |
| Dataset Phylogeny |
|---|
| Taxonomy Groups | Number of Scaffolds |
|---|
| Not Available | 2 |
| All Organisms → cellular organisms → Bacteria → Terrabacteria group → Chloroflexi | 1 |
| All Organisms → cellular organisms → Bacteria → Terrabacteria group → Chloroflexi → unclassified Chloroflexi → Chloroflexi bacterium | 1 |
| All Organisms → cellular organisms → Archaea → unclassified Archaea → archaeon | 1 |
| All Organisms → cellular organisms → Bacteria → PVC group → Planctomycetes → unclassified Planctomycetota → Planctomycetes bacterium B3_Pla | 1 |
Ecosystem and Geography
| Ecosystem Assignment (GOLD) |
|---|
| Name | Soil Microbial Communities From Risofladan, Vaasa, Finland |
| Type | Environmental |
| Taxonomy | Environmental → Terrestrial → Soil → Clay → Unclassified → Soil → Soil Microbial Communities From Risofladan, Vaasa, Finland |
| Alternative Ecosystem Assignments |
|---|
| Environment Ontology (ENVO) | terrestrial biome → land → soil |
| Earth Microbiome Project Ontology (EMPO) | Unclassified |
| Location Information |
|---|
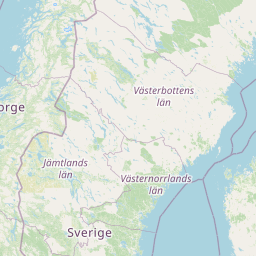
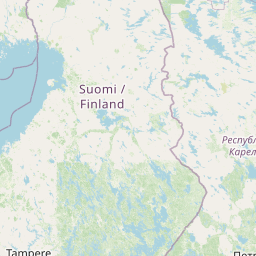
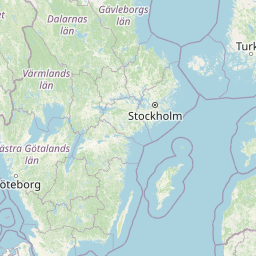
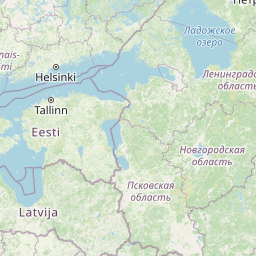
| Location | Finland: Risofladan, Vaasa |
| Coordinates | Lat. (o) | 63.0472 | Long. (o) | 21.7116 | Alt. (m) | N/A | Depth (m) | 1 |
| Location on Map |
|---|
|
|
| Zoom: |
Powered by OpenStreetMap © |
Associated Families
Associated Scaffolds
Sequences